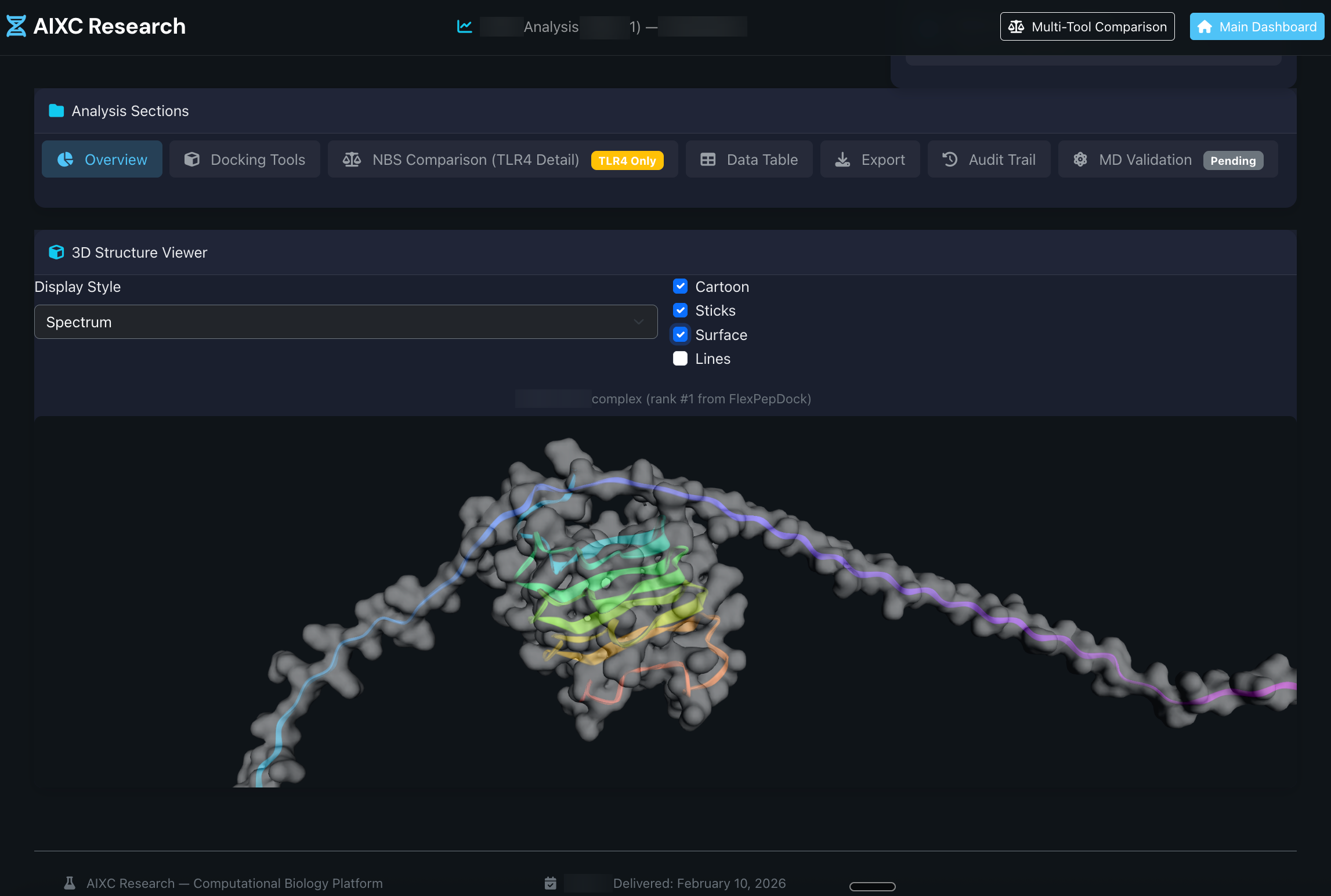
State-of-the-art protein structure prediction with Boltz-2our primary engine and three complementary backends. Template conditioning, multi-chain co-folding, and per-residue confidence metrics. 5-6x speedup.
Our primary engine is Boltz-2a state-of-the-art generative model, the latest generative structure prediction model. Three additional backends provide cross-validation and specialized capabilities.

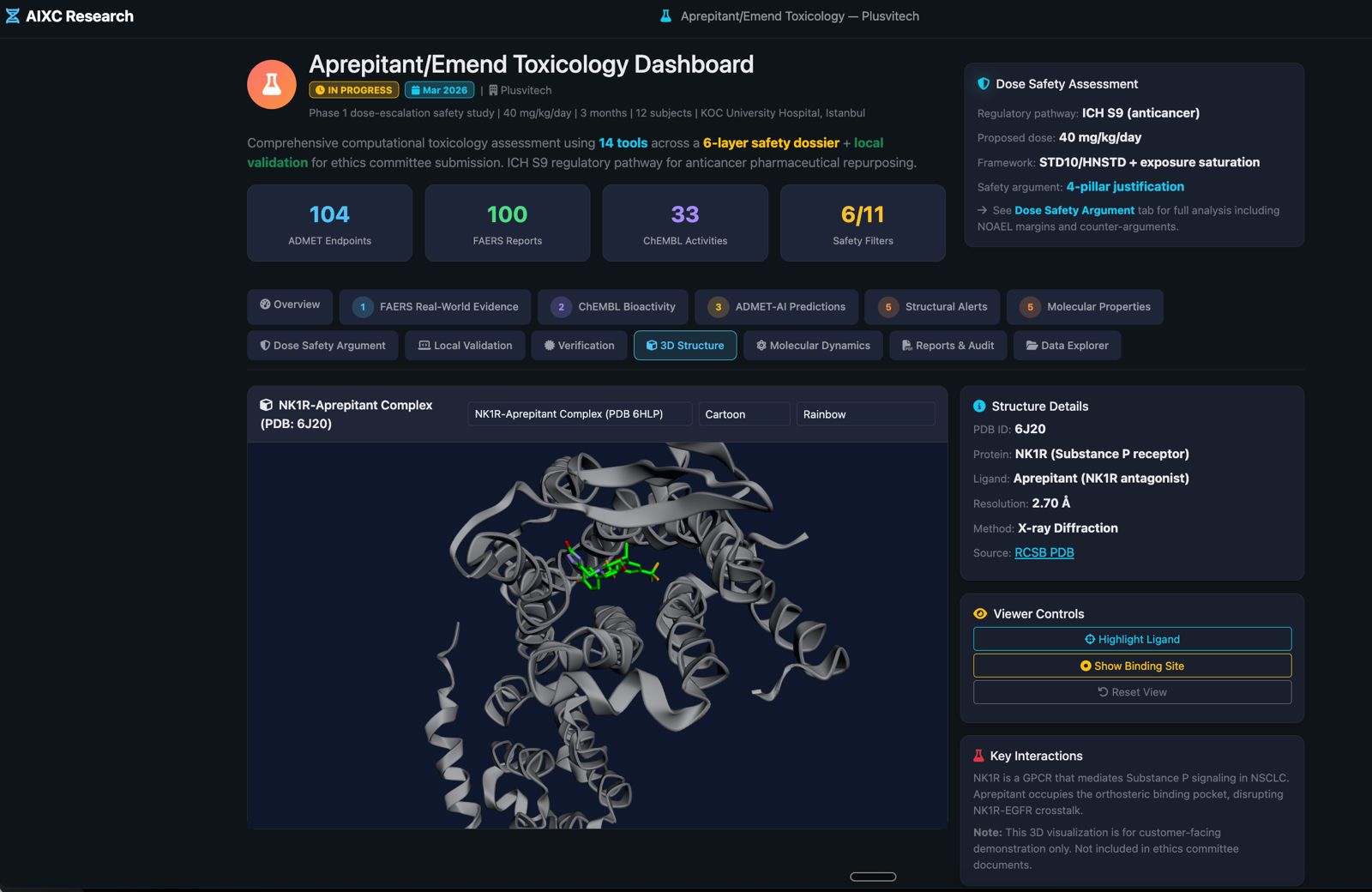
Beyond simple monomer prediction. Our pipeline handles the complex scenarios that real drug discovery projects require.
Every prediction comes with quantitative confidence scores. Know exactly which regions are reliable and which need experimental validation.
Our infrastructure runs on dedicated GPU clusters, delivering significant speedups over CPU-only inference.
How Boltz-2our primary engine compares to other leading structure prediction tools across key capabilities.
| Feature | AlphaFold2Method A | ESMFoldMethod B | Boltz-2AIXC Primary (AIXC) |
|---|---|---|---|
| Multi-chain co-folding | Limited | No | Full |
| Small molecule docking | No | No | Yes |
| Nucleic acid support | No | No | Yes |
| Template conditioning | Yes | No | Yes |
| MSA-free mode | No | Yes | Yes |
| Covalent modifications | No | No | Yes |
| Confidence metrics | pLDDT, pTM | pLDDT | pLDDT, pTM, PAE, ipTM |
| GPU support | No | CPU only | Full GPU |
Submit your protein sequence and we will deliver high-confidence 3D structure predictions with full confidence metrics and downloadable PDB/CIF files.